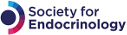Recent advances in genome editing are revolutionising the way we tackle big scientific questions. Tools have been developed that act like tiny ‘molecular scissors’, causing targeted genomic cuts in DNA. By exploiting cells’ innate DNA repair mechanisms, scientists can repair aberrant genes or modify the genetic code to their liking. With these tools, we can correct pathogenic mutations, model human genetic disease in cells and animal systems, modify key organisms for biotechnology and agriculture, and potentially eradicate hereditary monogenic disease.
RESTORING DNA BREAKS
A number of different methods exist to cause genomic cuts; however, repair mechanisms remain the same. DNA breaks can occur naturally through intracellular nucleases/reactive oxygen species or externally by ionising radiation/ultraviolet light. If left unrepaired, the damage will lead to cell death. Luckily, the cell has developed natural processes to fix breaks. Repair can follow one of two paths: non-homologous end joining (NHEJ) or homology-directed repair (HDR).1
NHEJ is a fast simple method, generally favoured when broken ends are compatible.2 Here, specific proteins guide the alignment of the broken ends, joining them together using DNA ligase IV.3 In the majority of cases, the correct ends rejoin. However, annealing incorrect ends or the removal of damaged nucleotides can lead to chromosomal aberrations and mutations.4 Hence, NHEJ is error prone, a characteristic researchers exploit to disrupt gene function.
Conversely, HDR uses an intact copy of the gene loci to repair the broken sequence, an accurate but slow method.2 After a DNA break event, pathways are activated to remove damaged nucleotides by nibbling forward and reverse DNA strands, leaving single-stranded DNA overhangs.5 Subsequently, the DNA ends are coated with recombinases and co-factors, forming homology-searching nucleoprotein filaments.6 The filaments hunt for sequence homologies in sister chromatids, which guide repair.7 Again, adept scientists have manipulated this process to knock-in genetic material to loci, repairing genetic variants or adding traceable tag proteins.8,9 Thus, DNA repair by NHEJ or HDR forms the basis of the genome editing technique.

A simplified schematic of CRISPR/Cas9-directed genome editing. (a) CRISPR/Cas9 complex is used to target genomic DNA breaks, like ‘tiny molecular scissors’. (b) DNA breaks can be repaired by either nonhomologous end joining (NHEJ, left) or homology-directed repair (HDR, right). (c) NHEJ is error prone, resulting in mutations in targeted genes, permitting gene function experiments. HDR can be used to replace genetic information, mimicking or correcting pathogenic sequences (e.g. for disease model
MAKING PRECISE CUTS
Genomic breaks caused by natural forces are unpredictable and occur randomly. To use DNA repair mechanisms for scientific gains, researchers have developed precision DNA cutting tools. Although other genome editing tools exist (meganucleases, zinc finger nucleases and transcription activator-like effector nucleases or TALENS10), CRISPR/Cas9 (clustered regularly interspaced short palindromic repeats with CRISPR associated protein 9) has proved to be efficient and accessible to researchers.
Coined as ‘the discovery of the century’, the two scientists attributed to its identification, Jennifer Doudna (Berkeley, CA, USA) and Emmanuelle Charpentier (Berlin, Germany),11 have been tipped to win a Nobel Prize for their work, along with Feng Zhang (Cambridge, MA, USA), a pioneer in the use of CRISPR/Cas9 on mammalian cells.12
First discovered as a bacterial adaptive immune response to foreign invading viral DNA,13 the system acts to incorporate specific portions of phage DNA into the CRISPR loci. This allows generation of homology-searching guide RNAs that bind with Cas9 endonuclease and target cuts in the viral DNA.13 Synthetic guide RNAs can be designed to incorporate user-defined target sequences, and synthesised in the laboratory. Therefore, the CRISPR/Cas9 system offers a versatile and adaptive tool for generating targeted genomic breaks.
A TOOLKIT TO TREAT DISEASE
CRISPR/Cas9 is a powerful tool and has been used to remove an erroneous exon from the dystrophin gene in mice with muscular dystrophy, resulting in restored muscle integrity and function.14–16 Furthermore, proof of principle experiments in human embryos (not destined for transplantation) used CRISPR/Cas9 and synthetic DNA template-mediated HDR to correct a mutation in MYBPC3, which normally causes sudden death syndrome.9 Despite the obvious ethical issues surrounding the misuse of genome editing to manipulate human genetics, there are serious concerns regarding inherited off-target effects that must be resolved before reaching the clinic.
DISSECTING THE ZEBRAFISH GENOME
Although genome editing continues to raise concerns for clinical use, it has provided immediate advances in modelling disease in animals. The zebrafish is one such organism that has benefited from improved genome editing techniques. Popular for its versatility as an in vivo model, the zebrafish provides rapid ex utero development, large numbers of embryos, ease of genetic and pharmaceutical manipulation, and over 82% disease-causing genes in common with humans.17 For many years, antisense morpholino (MO) technology has been used to knockdown gene function in zebrafish, by inhibiting gene-specific translation. However, MO use has come under scrutiny in recent years.18 Mutant zebrafish lines remain the ‘gold standard’, although generating them in the past required laborious large scale teratogen-based screens.19,20 Although many mutants have been recovered for a large number of genes, many more have been missed.
The targeted nature of CRISPR/Cas9 has meant that zebrafish mutations in genes known, or suspected, to be pathogenic can be easily disrupted as a consequence of error prone NHEJ.21 Although it is possible to create exact mutations that recapitulate patient variants by the addition of synthetic DNA templates for HDR, this is more challenging.22 However, the technology is constantly evolving; variations in endonuclease activity are being developed to improve specificity and efficiency.23
These are exciting times. Genome editing has become commonplace in most research laboratories and the possibilities for manipulating genomic DNA for scientific advances are endless.
Daniel Osborn, Senior Lecturer in Genetics, Genetics Research Centre at St George’s, University of London
REFERENCES
- Symington LS & Gautier J 2011 Annual Review of Genetics 45 247–271.
- Mao Z et al. 2008 DNA Repair 7 1765–1771.
- Lieber MR 2010 Annual Review of Biochemistry 79 181–211.
- Varga T & Aplan PD 2005 DNA Repair 4 1038–1046.
- Liu T & Huang J 2016 Genomics Proteomics & Bioinformatics 14 126–130.
- Krogh BO & Symington LS 2004 Annual Review of Genetics 38 233–271.
- Liang F et al. 1998 Proceedings of the National Academy of Sciences of the U S A 95 5172–5177.
- Ratz M et al. 2015 Scientific Reports 5 9592.
- Osborn DP et al. 2017 American Journal of Human Genetics 100 537–545.
- Guha TK et al. 2017 Computational & Structural Biotechnology Journal 15 146–160.
- Jinek M et al. 2012 Science 337 816–821.
- Cong L et al. 2013 Science 339 819–823.
- Horvath P & Barrangou R 2010 Science 327 167–170.
- Nelson CE et al. 2016 Science 351 403–407.
- Long C et al. 2016 Science 351 400–403.
- Tabebordbar M et al. 2016 Science 351 407–411.
- Howe K et al. 2013 Nature 496 498–503.
- Nasevicius A & Ekker SC 2000 Nature Genetics 26 216–220.
- Driever W et al. 1996 Development 123 37–46.
- Haffter P et al. 1996 Development 123 1–36.
- Talbot JC & Amacher SL 2014 Zebrafish 11 583–585.
- Albadri S et al. 2017 Methods 121–122 77–85.
- Richter F et al. 2017 Current Opinion in Biotechnology 48 119–126.