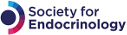A virtuous cycle in which a model trained against wet-lab data (top) is used to create predictions that lead to wet-lab tests (right) yielding data (bottom) that can be used to improve the model (left). Illustration by Kathryn Garner
Gonadotrophin-releasing hormone (GnRH) is a peptide hormone that is secreted from the hypothalamus in pulses to control reproduction. It does this via GnRH receptors (GnRH-Rs) on pituitary gonadotrophs that mediate its effects on the synthesis and secretion of luteinising hormone (LH) and follicle-stimulating hormone (FSH).
The stimulation paradigm is crucial, since pulsatile and sustained GnRH have very different effects on the target cells, and this phenomenon is exploited in therapeutic manipulation of the system.1 Understanding how the complex inputs are sensed and processed by gonadotrophs forms the core of our research, and has led us to use mathematical approaches to enrich our wet-lab data.
EARLY INSIGHTS INTO GnRH
The structural determination and synthesis of GnRH were landmarks in the development of the field of neuroendocrinology (for which the Nobel Prize was awarded in 1977). They were rapidly followed by attempts to understand its mechanism of action. In the early days, all that was really known was that GnRH acted via receptors to increase production of second messengers.
At the time, the logic of cell signalling was very simple: hypothesising that A→B→C, one would set to work with pharmacological inhibitors and mimics to prove or disprove the hypothesis. However, the original signalling pathway idea has given way to the notion that GnRH (like other extracellular signals) engages a complex signalling network with numerous sites of divergence and convergence, as well as routes for positive and negative feedback. We now work with increasingly complex network cartoons for the GnRH signalling architecture, yet these are still vast over-simplifications, not least because they largely ignore dynamics.
MATHEMATICAL REASONING
Several years ago, Craig was discussing how such a network might decode input dynamics with Krasimira Tsaneva-Atanasova, a colleague in mathematical engineering. On seeing our latest network topology model, she pointed out that it was far too complex to understand by ‘intuition’ alone.
She outlined a mechanistic modelling approach. We then set about developing a model for GnRH signalling based on a series of ordinary differential equations (ODEs) describing receptor occupancy through to G-protein-mediated activation of phospholipase C, calcium mobilisation, activation of the calcium-sensitive transcription factor NFAT (nuclear factor of activated T cells) and protein kinase C-mediated activation of ERK (extracellular signal regulated kinase).
‘Our modelling offers an entrée into a virtuous cycle in which a model trained against wet-lab data is used to create predictions that lead to wet-lab tests yielding data that can be used to improve the model, so that the improved model can be used for further predictions’
We included only molecules that have actually been shown to be necessary for GnRH action in vivo (GnRH, GnRH-R, ERK, gonadotrophin α-subunit, LHβ and FSHβ), with the addition of a calcium effector. Moreover, since ERK2-GFP (green fluorescent protein) and NFAT-EFP (emerald fluorescent protein) reporters both translocate from the cytoplasm to the nucleus on activation, we were able to use these as live cell read-outs for GnRH signalling during trains of GnRH pulses.2,3
Leaning heavily on previous GnRH signalling models, together with models for ERK activation, calcium mobilisation and NFAT activation, we developed a mechanistic model that was trained on and validated against our wet-lab data. This model has gone through multiple iterations. The earliest did not include GnRH-R trafficking and was trained on data for GnRH-R signalling in a HeLa cell model,4 whereas the latest incorporates GnRH-R internalisation and recycling and is trained against wet-lab data for signalling of endogenous GnRH-R in a gonadotroph-derived cell line.5
ENHANCED PREDICTIONS
It might seem that we are merely generating a mathematical description of what we’ve already seen in our wet-lab data. However, our modelling offers an entrée into a virtuous cycle in which a model trained against wet-lab data is used to create predictions that lead to wet-lab tests yielding data that can be used to improve the model, so that the improved model can be used for further predictions.
Wet-lab experiments with pulsatile GnRH are very labour intensive. Ideally, we’d like to vary pulse frequency, width and concentration in parallel, and this is straightforward in silico, but not practical in vitro. Instead, we use simulations to explore possible system behaviours and help us plan experiments to reveal actual behaviours.
ANSWERING QUESTIONS
In a recent study, we used a minimal model to address the fundamental question of why GnRH secretion is pulsatile. We found that the use of a pulsatile input could increase efficiency, generate specificity and make the system resistant to receptor trafficking.5 We also asked whether upstream negative feedback (i.e. receptor internalisation) could explain why maximal responses to GnRH are very often obtained with sub-maximal GnRH pulse frequencies. The simulations revealed that it could, but only under conditions where receptor internalisation rates are high and desensitisation of downstream responses are marked, neither of which are consistent with wet-lab data. Consequently, our current emphasis is on downstream (i.e. transcriptional) mechanisms that could achieve this.
Notably, our ODE approach models a single cell, yet in our wet-lab data we observe enormous cell–cell variability in signalling responses. We have addressed this by applying a statistical measure known as mutual information, originally developed to analyse electronic communications, to quantify the information transferred along our noisy signalling pathways.
In another recent study, we used this approach to consider the effects of negative feedback on sensing of GnRH, focusing on ERK-dependent feedback (rapid transcription-independent and slow transcription-dependent feedback) and on receptor desensitisation, finding that downstream adaptive ERK-mediated feedback fine-tunes hormone sensing, with optimal GnRH sensing at intermediate feedback intensities.6
Indeed, it is clear that we were unable to fully comprehend the GnRH network topology using our intuition alone. Beginning with mechanistic modelling, we have begun to unravel the complexities of pulsatile GnRH signalling and now consider the contribution of individual cells to the variability within a whole cell population.
Kathryn Garner & Craig McArdle
Laboratories for Integrative Neuroscience and Endocrinology, School of Clinical Sciences, University of Bristol
REFERENCES
- McArdle CA & Robertson MS 2015 In Knobil and Neill’s Physiology of Reproduction, Eds TM Plant & AJ Zeleznik, pp 335–397. Amsterdam: Elsevier.
- Armstrong SP et al. 2009 Journal of Biological Chemistry 284 35746–35757.
- Armstrong SP et al. 2010 Journal of Biological Chemistry 285 24360–24371.
- Tsaneva-Atanasova K et al. 2011 Journal of the Royal Society Interface 9 170–182.
- Pratap A et al. 2016 Molecular & Cellular Endocrinology doi:10.1016/ j.mce.2016.08.022.
- Garner KL et al. 2016 Journal of Biological Chemistry 291 2246–2259.